Overview
birddog helps you detect emergence and trace
trajectories in scientific literature and patents. It reads datasets
from OpenAlex and Web of Science (WoS), builds citation-based networks,
identifies groups, and summarizes their dynamics.
The stable release is on CRAN. The development version is available on GitHub: https://github.com/roneyfraga/birddog.
Installation
# stable version (CRAN)
install.packages("birddog")
# development version (GitHub)
# install.packages("remotes")
remotes::install_github("roneyfraga/birddog")Data sources
birddog supports two main data sources:
- OpenAlex: browser search with CSV export, or API via openalexR.
-
Web of
Science: multiple export formats (
.bib,.ris, plain-text.txt, tab-delimited.txt).
OpenAlex via API or CSV
library(openalexR)
# Fetch works from OpenAlex API
url_api <- "https://api.openalex.org/works?page=1&filter=primary_location.source.id:s121026525"
openalexR::oa_request(query_url = url_api) |>
openalexR::oa2df(entity = "works") |>
birddog::read_openalex(format = "api") ->
M
# Or from a CSV export
M <- birddog::read_openalex("path/to/openalex-export.csv", format = "csv")Web of Science (WoS)
# BibTeX
M <- birddog::read_wos("path/to/savedrecs.bib", format = "bib")
# RIS
M <- birddog::read_wos("path/to/savedrecs.ris", format = "ris")
# Plain text
M <- birddog::read_wos("path/to/savedrecs.txt", format = "txt-plain-text")
# Tab-delimited
M <- birddog::read_wos("path/to/savedrecs.txt", format = "txt-tab-delimited")Example dataset
We use a biogas dataset from OpenAlex with 57,734 documents as a running example.
# Download from OpenAlex (~15 min)
query_oa <- "( biogas )"
openalexR::oa_fetch(
entity = "works",
title_and_abstract.search = query_oa,
verbose = TRUE
) ->
papers
M <- birddog::read_openalex(papers, format = "api")
# Pre-computed dataset
url_m <- "https://roneyfraga.com/volume/keep_it/biogas-data/M.rds"
M <- readRDS(url(url_m))
dplyr::glimpse(M)
#> Rows: 57,734
#> Columns: 55
#> $ id <chr> "https://openalex.org/W2072823483", "https…
#> $ id_short <chr> "W2072823483", "W2109587007", "W2032792259…
#> $ SR <chr> "W2072823483", "W2109587007", "W2032792259…
#> $ PY <int> 2009, 2008, 2009, 2011, 2018, 2015, 2006, …
#> $ TC <int> 2672, 2542, 1616, 1228, 1111, 1540, 667, 9…
#> $ TI <chr> "Biogas production: current state and pers…
#> $ DI <chr> "https://doi.org/10.1007/s00253-009-2246-7…
#> $ AB <chr> NA, "Lignocelluloses are often a major or …
#> $ CR <chr> "W1417235137;W1503353321;W1506760845;W1529…
#> $ DE <chr> "biogas;digestate;renewable natural gas", …
#> $ AU <chr> "PETER WEILAND", "MOHAMMAD J. TAHERZADEH;K…
#> $ DB <chr> "openalex_api", "openalex_api", "openalex_…
#> $ title <chr> "Biogas production: current state and pers…
#> $ display_name <chr> "Biogas production: current state and pers…
#> $ authorships <list> [<tbl_df[1 x 7]>], [<tbl_df[2 x 7]>], [<t…
#> $ abstract <chr> NA, "Lignocelluloses are often a major or …
#> $ doi <chr> "https://doi.org/10.1007/s00253-009-2246-7…
#> $ publication_date <date> 2009-09-23, 2008-09-01, 2009-02-14, 2011-…
#> $ publication_year <int> 2009, 2008, 2009, 2011, 2018, 2015, 2006, …
#> $ relevance_score <dbl> 1861.7662, 1781.5554, 1455.7186, 1270.2125…
#> $ fwci <dbl> 12.110, 6.397, 17.615, 22.271, 68.880, 115…
#> $ cited_by_count <int> 2672, 2542, 1616, 1228, 1111, 1540, 667, 9…
#> $ counts_by_year <list> [<data.frame[14 x 2]>], [<data.frame[14 x…
#> $ cited_by_api_url <chr> "https://api.openalex.org/works?filter=cit…
#> $ ids <list> <"https://openalex.org/W2072823483", "htt…
#> $ type <chr> "review", "review", "review", "article", "…
#> $ is_oa <lgl> FALSE, TRUE, FALSE, FALSE, TRUE, FALSE, FA…
#> $ is_oa_anywhere <lgl> FALSE, TRUE, FALSE, FALSE, TRUE, FALSE, FA…
#> $ oa_status <chr> "closed", "gold", "closed", "closed", "hyb…
#> $ oa_url <chr> NA, "https://www.mdpi.com/1422-0067/9/9/16…
#> $ any_repository_has_fulltext <lgl> FALSE, TRUE, FALSE, FALSE, FALSE, FALSE, F…
#> $ source_display_name <chr> "Applied Microbiology and Biotechnology", …
#> $ source_id <chr> "https://openalex.org/S70979998", "https:/…
#> $ issn_l <chr> "0175-7598", "1422-0067", "0960-8524", "09…
#> $ host_organization <chr> "https://openalex.org/P4310319900", "https…
#> $ host_organization_name <chr> "Springer Science+Business Media", "Multid…
#> $ landing_page_url <chr> "https://doi.org/10.1007/s00253-009-2246-7…
#> $ pdf_url <chr> NA, "https://www.mdpi.com/1422-0067/9/9/16…
#> $ license <chr> NA, NA, NA, NA, "cc-by", NA, NA, NA, NA, N…
#> $ version <chr> NA, "publishedVersion", NA, NA, "published…
#> $ referenced_works <list> <"https://openalex.org/W1417235137", "htt…
#> $ referenced_works_count <int> 55, 193, 9, 10, 14, 145, 16, 0, 211, 50, 1…
#> $ related_works <list> <"https://openalex.org/W4387315092", "htt…
#> $ concepts <list> [<data.frame[21 x 5]>], [<data.frame[18 x…
#> $ topics <list> [<tbl_df[12 x 5]>], [<tbl_df[12 x 5]>], […
#> $ keywords <list> [<data.frame[3 x 3]>], [<data.frame[5 x 3…
#> $ is_paratext <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, …
#> $ is_retracted <lgl> FALSE, FALSE, FALSE, FALSE, FALSE, FALSE, …
#> $ language <chr> "en", "en", "en", "en", "en", "en", "en", …
#> $ grants <list> NA, NA, NA, NA, NA, NA, NA, NA, NA, NA, <…
#> $ apc <list> [<data.frame[2 x 5]>], [<data.frame[2 x 5…
#> $ first_page <chr> "849", "1621", "5478", "1633", "457", "540…
#> $ last_page <chr> "860", "1651", "5484", "1645", "472", "555…
#> $ volume <chr> "85", "9", "100", "35", "129", "45", "32",…
#> $ issue <chr> "4", "9", "22", "5", NA, NA, "8", "05", NA…Citation network
Build a citation network to map the relationships between documents. Direct citation captures time-ordered influence; bibliographic coupling groups papers that share references.
net <- birddog::sniff_network(M, type = "direct citation")
net |>
tidygraph::activate(nodes) |>
dplyr::select(name, AU, PY, TI, TC) |>
dplyr::arrange(dplyr::desc(TC))
#> # A tbl_graph: 29490 nodes and 197059 edges
#> #
#> # A directed simple graph with 297 components
#> #
#> # Node Data: 29,490 × 5 (active)
#> name AU PY TI TC
#> <chr> <chr> <int> <chr> <int>
#> 1 W2072823483 PETER WEILAND 2009 Biog… 2672
#> 2 W2109587007 MOHAMMAD J. TAHERZADEH;KEIKHOSRO KARIMI 2008 Pret… 2542
#> 3 W2125363721 DAVID SCHMEIDLER 1969 The … 1873
#> 4 W2089274821 İRINI ANGELIDAKI;M. M. ALVES;DAVID BOLZONELLA;… 2009 Defi… 1857
#> 5 W2075740579 PERRY L. MCCARTY;JAEHO BAE;JEONGHWAN KIM 2011 Dome… 1670
#> 6 W2032792259 JENS BO HOLM‐NIELSEN;TEODORITA AL SEADI;PIOTR … 2009 The … 1616
#> 7 W2024397118 CHUNLAN MAO;YONGZHONG FENG;XIAOJIAO WANG;GUANG… 2015 Revi… 1540
#> 8 W2043402970 YI ZHENG;JIA ZHAO;FUQING XU;YEBO LI 2014 Pret… 1238
#> 9 W2018893323 ELINE RYCKEBOSCH;MARGRIET DROUILLON;HAN VERVAE… 2011 Tech… 1228
#> 10 W2111419835 CUNSHENG ZHANG;HAIJIA SU;JAN BAEYENS;TIANWEI T… 2014 Revi… 1173
#> # ℹ 29,480 more rows
#> #
#> # Edge Data: 197,059 × 2
#> from to
#> <int> <int>
#> 1 1 48
#> 2 1 80
#> 3 1 77
#> # ℹ 197,056 more rowsComponents
Identify connected components to eliminate disconnected documents that do not share the same bibliographic references.
comps <- birddog::sniff_components(net)
comps$components |>
dplyr::slice_head(n = 5) |>
gt::gt()| component | quantity_publications | average_age |
|---|---|---|
| c1 | 28820 | 2016.840 |
| c2 | 27 | 2018.444 |
| c3 | 7 | 2014.857 |
| c4 | 7 | 2011.857 |
| c5 | 7 | 2019.286 |
Groups (community detection)
Detect research communities within the citation network. Each group represents a cluster of related publications.
groups <- birddog::sniff_groups(
comps,
algorithm = "fast_greedy",
min_group_size = 30,
seed = 888L
)
groups$aggregate |>
gt::gt()| group | quantity_papers | average_age |
|---|---|---|
| c1g1 | 6448 | 2017.027 |
| c1g2 | 4157 | 2015.351 |
| c1g3 | 2960 | 2016.728 |
| c1g4 | 2893 | 2017.972 |
| c1g5 | 1937 | 2017.712 |
| c1g6 | 1805 | 2015.425 |
| c1g7 | 1588 | 2017.868 |
| c1g8 | 1529 | 2017.650 |
| c1g9 | 1289 | 2016.700 |
| c1g10 | 924 | 2017.745 |
| c1g11 | 853 | 2017.613 |
| c1g12 | 818 | 2017.555 |
| c1g13 | 598 | 2015.572 |
| c1g14 | 442 | 2017.713 |
| c1g15 | 295 | 2011.854 |
| c1g16 | 47 | 2013.319 |
| c1g17 | 31 | 2019.548 |
Group attributes
Summarize group-level statistics including publication trends and growth rates.
# ~2 min
groups_attributes <- birddog::sniff_groups_attributes(
groups,
growth_rate_period = 2010:2024,
show_results = FALSE
)
groups_attributes$attributes_table| Groups Attributes | |||||
| Group | Publications | Average age1 | Growth rate2 | Doubling time3 | Horizon plot4 |
|---|---|---|---|---|---|
| c1g1 | 6448 | 2017+0m | 6.5 | 11y+1m | |
| c1g2 | 4157 | 2015+4m | 4.9 | 15y+7m | |
| c1g3 | 2960 | 2016+9m | 3.7 | 19y+2m | |
| c1g4 | 2893 | 2017+12m | 9.0 | 8y | |
| c1g5 | 1937 | 2017+9m | 10.0 | 7y+4m | |
| c1g6 | 1805 | 2015+5m | 2.9 | 24y+6m | |
| c1g7 | 1588 | 2017+10m | 8.6 | 8y+5m | |
| c1g8 | 1529 | 2017+8m | 11.2 | 7y+7m | |
| c1g9 | 1289 | 2016+8m | 6.0 | 12y+11m | |
| c1g10 | 924 | 2017+9m | 7.1 | 10y+1m | |
| c1g11 | 853 | 2017+7m | 6.9 | 10y+5m | |
| c1g12 | 818 | 2017+7m | 8.4 | 9y+7m | |
| c1g13 | 598 | 2015+7m | 4.6 | 15y+4m | |
| c1g14 | 442 | 2017+9m | 11.3 | 6y+6m | |
| c1g15 | 295 | 2011+10m | 4.8 | 15y+8m | |
| c1g16 | 47 | 2013+4m | 1.3 | 54y+6m | |
| c1g17 | 31 | 2019+7m | 16.3 | 5y+7m | |
| 1 Average publication year: For example, '2016+7m' means that the articles were published, on average, in 2016 plus seven months. | |||||
| 2 Growth rate percentage year. Calculated by exp(b1)-1 where b1 is the econometric model coefficient. Time span, 2010 until 2024. | |||||
| 3 y = years, m = months. Calculated by ln(2)/b1 where b1 is the econometric model coefficient. | |||||
| 4 Publications between 2010 and 2024. Chart type horizon plot. | |||||
| Source: OpenAlex. Data extracted, organized and estimated by the authors. | |||||
Group keywords
Explore the most frequent keywords in each group.
groups_keywords <- birddog::sniff_groups_keywords(groups)
groups_keywords |>
dplyr::filter(group %in% c('c1g1', 'c1g2', 'c1g3')) |>
gt::gt()| group | term_freq | term_tfidf |
|---|---|---|
| c1g1 | biogas (5702); biogas production (2089); digestion (942); mesophile (785); food waste (644); biodegradable waste (563); cow dung (515); sewage sludge (498); hydraulic retention time (336); digestate (308); lignocellulosic biomass (261); total dissolved solids (228); hemicellulose (203); (188); hyacinth (168) | food waste [0.004] (644); gompertz function [0.004] (150); hyacinth [0.0036] (168); lignocellulosic biomass [0.0035] (261); steam explosion [0.003] (69); total dissolved solids [0.003] (228); digestion [0.0028] (942); hemicellulose [0.0027] (203); mesophile [0.0024] (785); enzymatic hydrolysis [0.0023] (88); sodium hydroxide [0.0023] (60); corn stover [0.0022] (82); stalk [0.0022] (49); sonication [0.0021] (55); eichhornia crassipes [0.002] (76) |
| c1g2 | biogas (3485); biogas production (847); cow dung (261); (212); biodegradable waste (193); energy source (156); animal waste (146); renewable resource (134); mesophile (120); firewood (109); digestion (102); digestate (99); stove (95); food waste (93); investment (85) | firewood [0.0069] (109); stove [0.0051] (95); diesel generator [0.0048] (46); emergy [0.0048] (33); rural electrification [0.0046] (44); dome (geology) [0.0045] (31); microgrid [0.0044] (50); kerosene [0.0036] (41); liquefied petroleum gas [0.0036] (56); hybrid power [0.0035] (34); energy poverty [0.003] (14); payback period [0.0029] (82); deforestation [0.0028] (27); promotion (chess) [0.0025] (24); consumption [0.0024] (46) |
| c1g3 | biogas (2368); biogas production (588); energy crop (281); silage (221); (217); digestate (213); food waste (97); biodegradable waste (87); cogeneration (84); renewable resource (75); digestion (68); manure management (66); investment (65); hectare (58); sewage sludge (58) | hectare [0.0072] (58); energy crop [0.0049] (281); sweet sorghum [0.0044] (22); arable land [0.0043] (41); silage [0.0039] (221); phalaris arundinacea [0.0033] (11); agricultural land [0.0031] (13); arundo donax [0.0027] (11); manure management [0.0025] (66); crop rotation [0.0024] (12); externality [0.0024] (10); supply chain optimization [0.0024] (10); fodder [0.0024] (19); triticale [0.0023] (22); cogeneration [0.0023] (84) |
Group NLP terms
Extract key phrases from abstracts using natural language processing.
# ~30 min
groups_terms <- birddog::sniff_groups_terms(groups, algorithm = "phrase")
groups_terms$terms_table |>
dplyr::slice_head(n = 3) |>
gt::gt()| group | term_freq | term_tfidf |
|---|---|---|
| c1g1 | biogas production (6713); anaerobic digestion (5247); methane production (1419); food waste (1344); anaerobic co-digestion (1005); methane yield (922); biogas yield (879); cow dung (818); solid waste (804); sewage sludge (759); volatile solids (668); renewable energy (638); water hyacinth (633); retention time (624); digestion process (567) | water hyacinth [0.0017] (633); anaerobic digestion [0.0016] (5247); thermal pretreatment [0.0013] (123); wheat straw [0.0010] (382); anaerobic co-digestion [0.0010] (1005); hydrodynamic cavitation [0.0009] (66); gompertz model [0.0009] (174); coffee pulp [0.0009] (82); production from rice [0.0008] (59); naoh pretreatment [0.0008] (58); sugarcane bagasse [0.0008] (132); mixing ratios [0.0008] (107); rumen fluid [0.0008] (147); orange peel [0.0007] (53); crude glycerol [0.0007] (84) |
| c1g2 | biogas production (1747); renewable energy (1348); anaerobic digestion (988); biogas plants (726); biogas plant (654); biogas technology (638); cow dung (572); energy sources (512); rural areas (388); organic waste (360); case study (343); biogas digester (331); solid waste (328); waste management (318); energy source (290) | technology adoption [0.0022] (83); rural households [0.0021] (102); household biogas [0.0020] (241); domestic biogas [0.0018] (91); biogas digesters [0.0018] (253); cassava peels [0.0018] (87); biogas technology adoption [0.0017] (62); household energy [0.0016] (81); biogas technology [0.0016] (638); rural biogas [0.0015] (73); rural household [0.0014] (84); biogas adoption [0.0014] (51); hybrid system [0.0013] (115); adoption of biogas [0.0013] (81); adoption of biogas technology [0.0013] (47) |
| c1g3 | biogas production (1711); anaerobic digestion (1040); biogas plants (1010); renewable energy (656); biogas plant (638); life cycle (420); greenhouse gas (351); agricultural biogas (321); energy production (301); cycle assessment (300); life cycle assessment (274); energy crops (266); energy sources (261); case study (247); dry matter (217) | cross ref [0.0072] (191); lt ;w [0.0053] (140); lt ;w :lsdexception [0.0041] (109); ;w :lsdexception [0.0041] (109); 2021 cross [0.0031] (82); cup plant [0.0030] (78); 2020 cross [0.0025] (67); agricultural biogas plants [0.0021] (151); 2022 cross [0.0021] (55); operations research [0.0021] (55); silage maize [0.0020] (69); 2019 cross [0.0019] (50); 2021 cross ref [0.0018] (47); energy crops [0.0015] (266); agricultural biogas [0.0015] (321) |
Hubs
Classify documents by their role in the network using the Zi-Pi method. Hub documents connect different research communities.
# ~20 min
groups_hubs <- birddog::sniff_groups_hubs(groups)
groups_hubs |>
dplyr::filter(zone != "noHub") |>
dplyr::mutate(Zi = round(Zi, 2), Pi = round(Pi, 2)) |>
dplyr::arrange(dplyr::desc(zone), dplyr::desc(Zi)) |>
dplyr::slice_head(n = 15) |>
gt::gt() |>
gt::text_transform(
locations = gt::cells_body(columns = name),
fn = function(x) {
glue::glue('<a href="https://openalex.org/{x}" target="_blank">{x}</a>')
}
)| group | name | TC | Ki | ki | Zi | Pi | zone |
|---|---|---|---|---|---|---|---|
| c1g1 | W2024397118 | 1540 | 524 | 243 | 17.45 | 0.75 | R7 |
| c1g6 | W2072823483 | 2672 | 1101 | 217 | 17.20 | 0.87 | R7 |
| c1g3 | W1972749747 | 737 | 344 | 138 | 13.67 | 0.78 | R7 |
| c1g3 | W2794649776 | 1111 | 523 | 127 | 12.54 | 0.86 | R7 |
| c1g1 | W1606100078 | 939 | 410 | 154 | 10.90 | 0.81 | R7 |
| c1g5 | W2032792259 | 1616 | 551 | 97 | 9.03 | 0.88 | R7 |
| c1g13 | W1985829371 | 227 | 89 | 34 | 8.20 | 0.78 | R7 |
| c1g1 | W2730727434 | 527 | 208 | 79 | 5.38 | 0.79 | R7 |
| c1g12 | W2029511769 | 308 | 96 | 34 | 5.04 | 0.78 | R7 |
| c1g13 | W2338842862 | 132 | 50 | 21 | 4.83 | 0.78 | R7 |
| c1g1 | W2049325764 | 385 | 163 | 64 | 4.27 | 0.79 | R7 |
| c1g5 | W2156879463 | 437 | 119 | 45 | 3.96 | 0.78 | R7 |
| c1g3 | W2798048710 | 329 | 160 | 43 | 3.93 | 0.82 | R7 |
| c1g1 | W2276289647 | 292 | 131 | 59 | 3.90 | 0.76 | R7 |
| c1g8 | W1912164921 | 488 | 132 | 34 | 3.82 | 0.85 | R7 |
Indexes: Citations Cycle Time
Measure the pace of change in each group by tracking how old the cited references are over time.
# ~1.5 min
groups_cct <- birddog::sniff_citations_cycle_time(
groups,
scope = "groups",
start_year = 2000,
end_year = 2024
)
groups_cct$plots[["c1g3"]]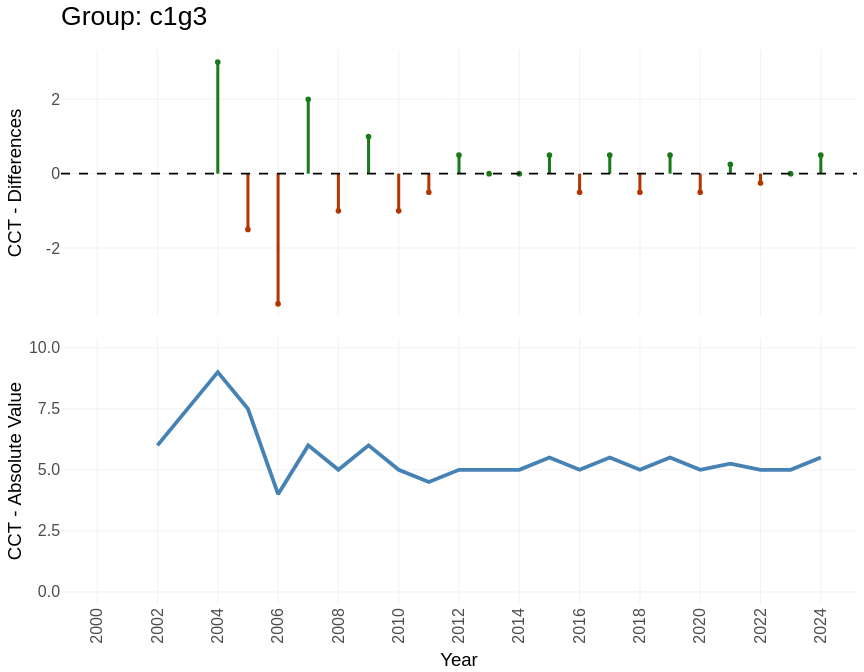
Indexes: Entropy
Track keyword diversity within each group over time. Increasing entropy signals thematic diversification; decreasing entropy signals convergence.
groups_entropy <- birddog::sniff_entropy(
groups,
scope = "groups",
start_year = 2000,
end_year = 2024
)
groups_entropy$plots[["c1g3"]]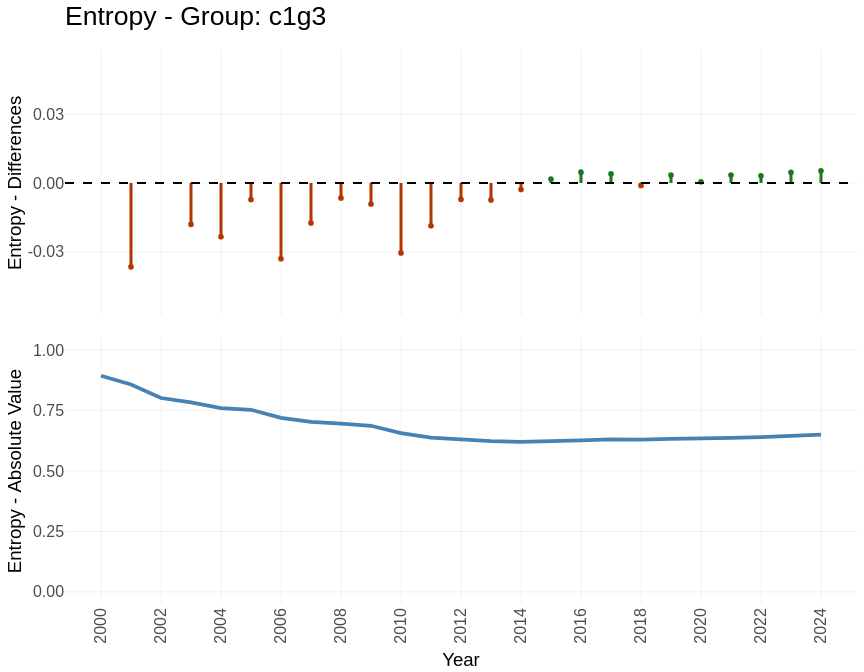
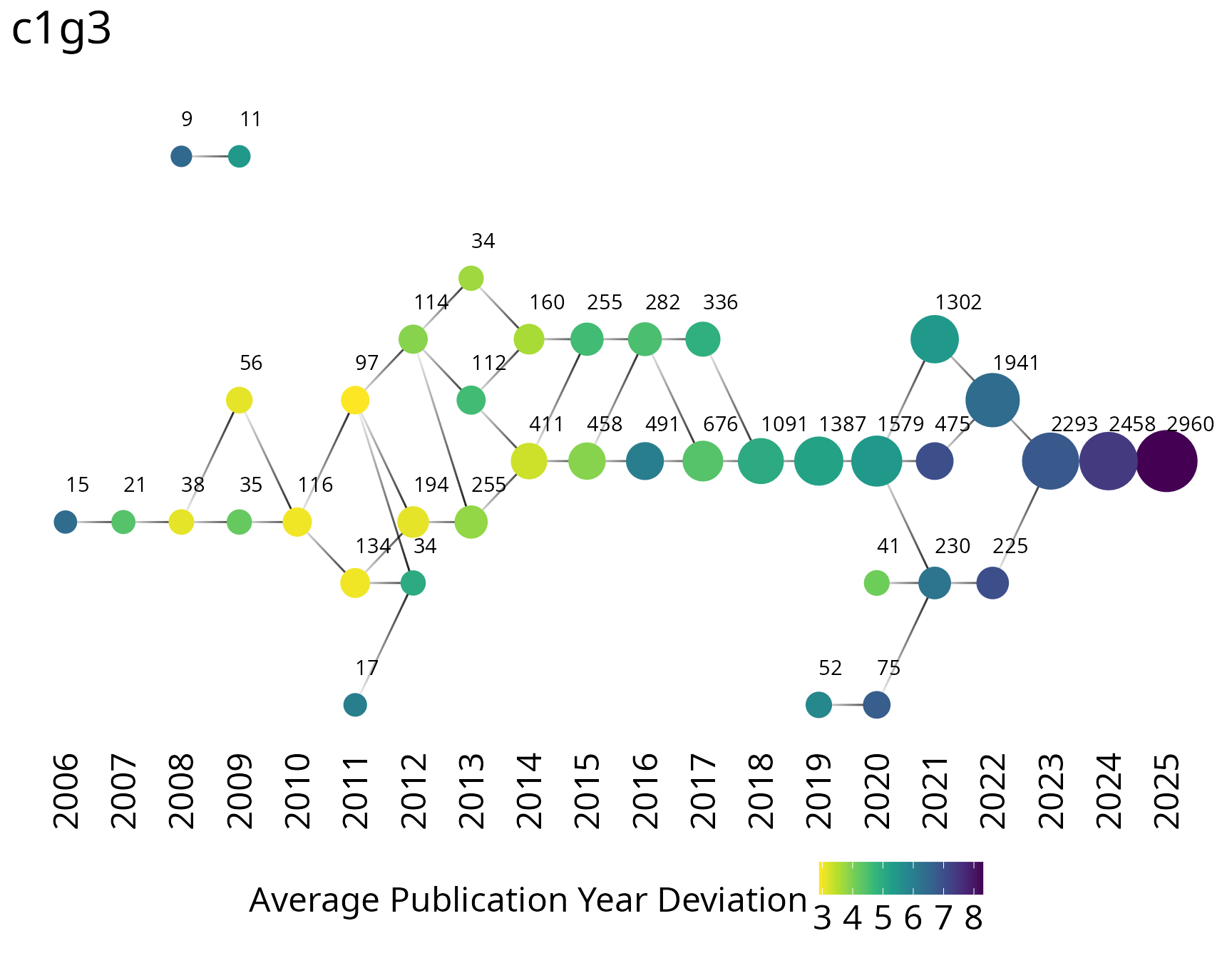
Group trajectories
Track how research communities evolve over time by building cumulative networks at each year and following groups through consecutive periods.
# ~2 min
groups_cumulative <- birddog::sniff_groups_cumulative(groups)
suppressMessages({
groups_cumulative_trajectories <- birddog::sniff_groups_trajectories(groups_cumulative)
})
birddog::plot_group_trajectories_2d(
groups_cumulative_trajectories,
group = "c1g3",
label_vertical_position = -2
)
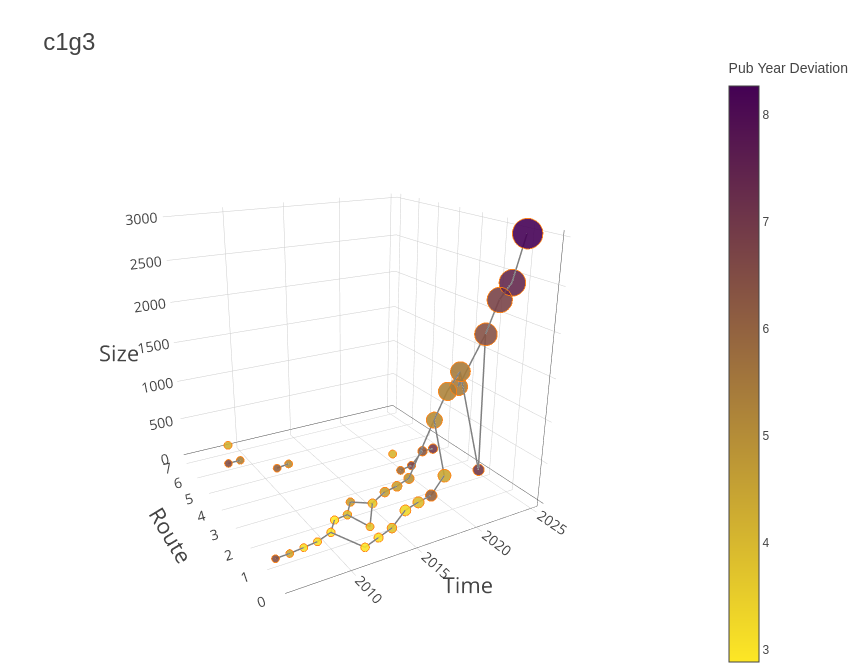
birddog::plot_group_trajectories_3d(
groups_cumulative_trajectories,
group = "c1g3"
)
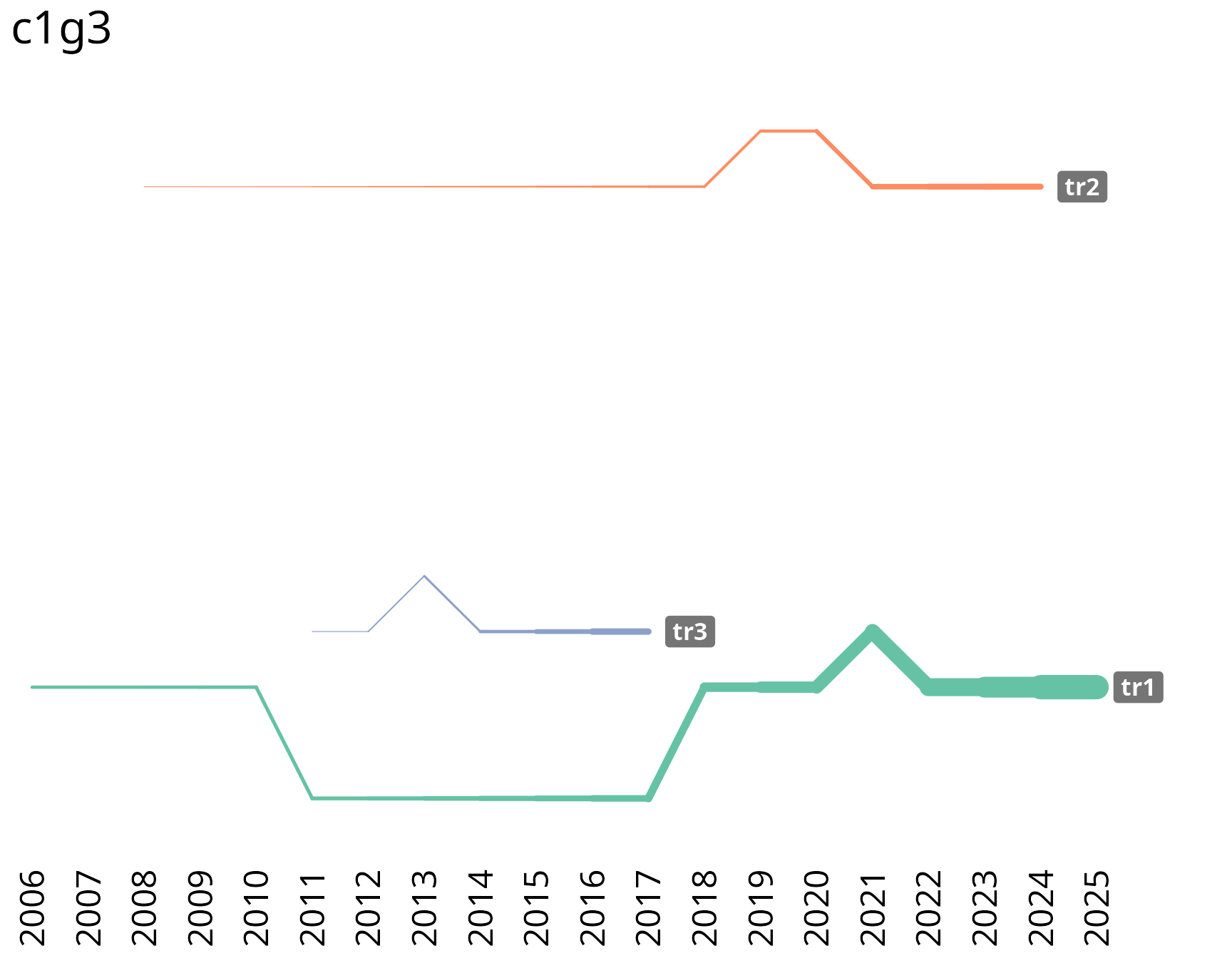
Trajectory detection and variable-width lines
Automatically detect the highest-scoring temporal paths within a group using dynamic programming, then display them as variable-width lines that grow with cumulative paper counts.
traj_data <- birddog::detect_main_trajectories(
groups_cumulative_trajectories,
group = "c1g3"
)
traj_filtered <- birddog::filter_trajectories(
traj_data$trajectories,
top_n = 3
)
birddog::plot_group_trajectories_lines_2d(
traj_data = traj_data,
traj_filtered = traj_filtered,
title = "c1g3"
)
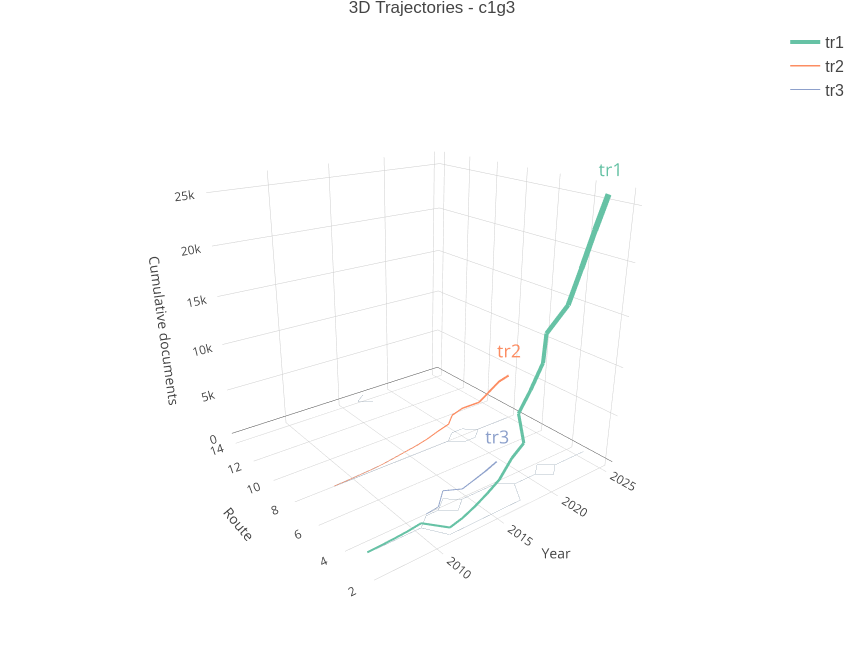
birddog::plot_group_trajectories_lines_3d(
traj_data = traj_data,
traj_filtered = traj_filtered,
group_id = "c1g3"
)
Citation growth per document
Track how individual documents accumulate citations over time to identify fast-growing papers.
# ~11 min
groups_cumulative_citations <- birddog::sniff_groups_cumulative_citations(
groups,
min_citations = 2
)Main Path Analysis
Identify the key route through the citation network, revealing the most influential chain of documents over time.
groups_key_route <- birddog::sniff_key_route(groups, scope = "groups")
groups_key_route[["c1g3"]]$plot
groups_key_route[["c1g3"]]$data |>
dplyr::select(-name) |>
gt::gt()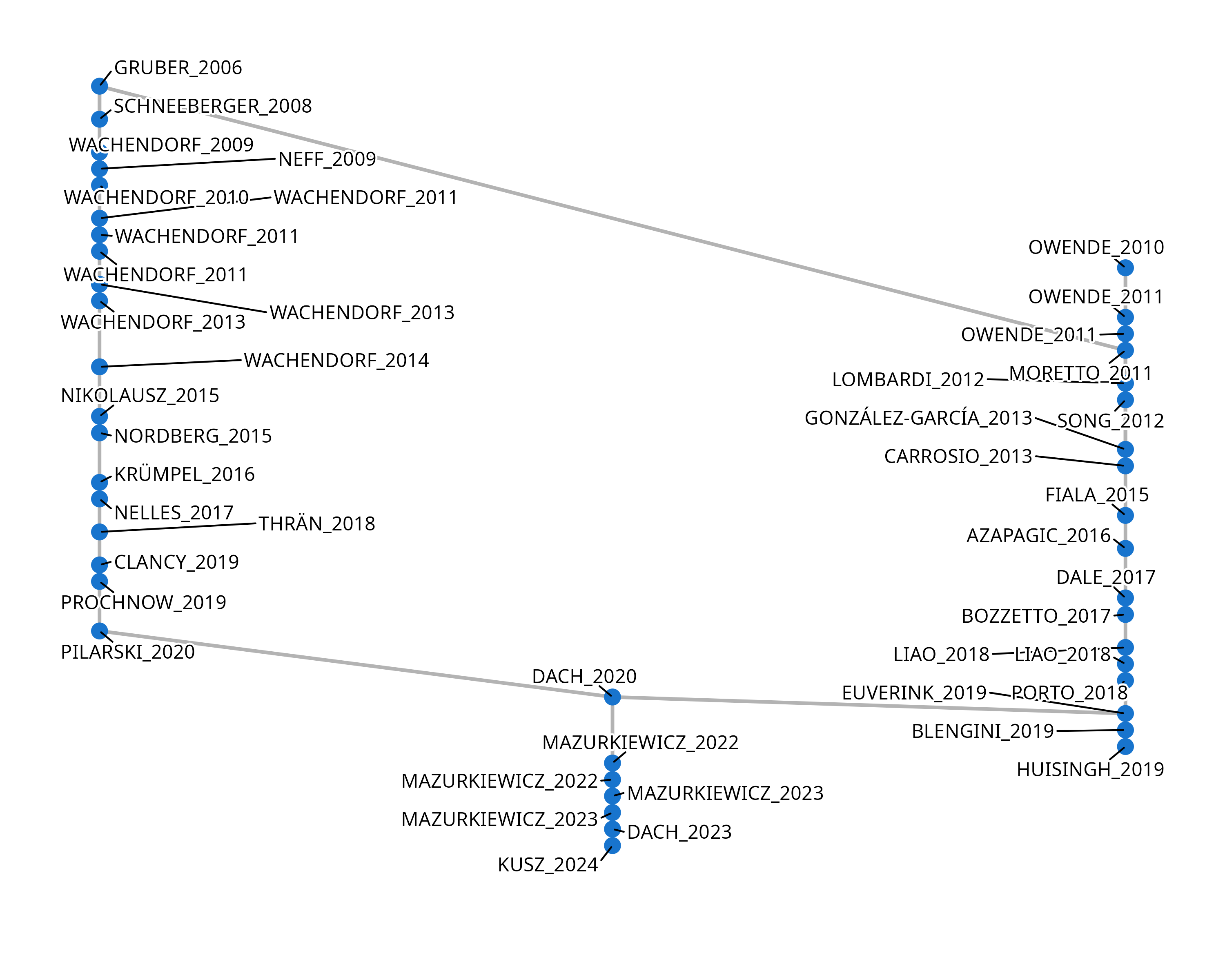
key_route_c1g3_data |>
dplyr::select(document = name, name2, title = TI) |>
gt::gt() |>
gt::text_transform(
locations = gt::cells_body(columns = document),
fn = function(x) {
glue::glue('<a href="https://openalex.org/{x}" target="_blank">{x}</a>')
}
)| document | name2 | title |
|---|---|---|
| W2008561700 | GRUBER_2006 | Biogas production from maize and dairy cattle manure—Influence of biomass composition on the methane yield |
| W2033886695 | SCHNEEBERGER_2008 | The optimal size for biogas plants |
| W2158862659 | NEFF_2009 | Utilization of semi‐natural grassland through integrated generation of solid fuel and biogas from biomass. I. Effects of hydrothermal conditioning and mechanical dehydration on mass flows of organic and mineral plant compounds, and nutrient balances |
| W2019905517 | WACHENDORF_2009 | Utilization of semi‐natural grassland through integrated generation of solid fuel and biogas from biomass. II. Effects of hydrothermal conditioning and mechanical dehydration on anaerobic digestion of press fluids |
| W1972749747 | OWENDE_2010 | Evaluation of energy efficiency of various biogas production and utilization pathways |
| W2134179273 | WACHENDORF_2010 | Utilization of semi‐natural grassland through integrated generation of solid fuel and biogas from biomass. III. Effects of hydrothermal conditioning and mechanical dehydration on solid fuel properties and on energy and greenhouse gas balances |
| W2003509580 | OWENDE_2011 | Environmental impacts of biogas deployment – Part II: life cycle assessment of multiple production and utilization pathways |
| W1997982003 | OWENDE_2011 | Environmental impacts of biogas deployment – Part I: life cycle inventory for evaluation of production process emissions to air |
| W2023450208 | WACHENDORF_2011 | Integrated generation of solid fuel and biogas from green cut material from landscape conservation and private households |
| W3124542086 | MORETTO_2011 | Investing in biogas: Timing, technological choice and the value of flexibility from input mix |
| W2075790500 | WACHENDORF_2011 | Influence of sward maturity and pre-conditioning temperature on the energy production from grass silage through the integrated generation of solid fuel and biogas from biomass (IFBB): 1. The fate of mineral compounds |
| W2091870505 | WACHENDORF_2011 | Influence of sward maturity and pre-conditioning temperature on the energy production from grass silage through the integrated generation of solid fuel and biogas from biomass (IFBB): 2. Properties of energy carriers and energy yield |
| W2078934777 | SONG_2012 | Life-cycle energy production and emissions mitigation by comprehensive biogas–digestate utilization |
| W2051151463 | LOMBARDI_2012 | Environmental analysis of biogas production systems |
| W2044886835 | WACHENDORF_2013 | Review of concepts for a demand-driven biogas supply for flexible power generation |
| W2064487381 | GONZÁLEZ‐GARCÍA_2013 | Anaerobic digestion of different feedstocks: Impact on energetic and environmental balances of biogas process |
| W2046271618 | CARROSIO_2013 | Energy production from biogas in the Italian countryside: Policies and organizational models |
| W2010747108 | WACHENDORF_2013 | Energetic conversion of European semi-natural grassland silages through the integrated generation of solid fuel and biogas from biomass: Energy yields and the fate of organic compounds |
| W2065522991 | WACHENDORF_2014 | Cost analysis of concepts for a demand oriented biogas supply for flexible power generation |
| W2191399137 | NIKOLAUSZ_2015 | Changing Feeding Regimes To Demonstrate Flexible Biogas Production: Effects on Process Performance, Microbial Community Structure, and Methanogenesis Pathways |
| W2048561114 | NORDBERG_2015 | Demand-Orientated Power Production from Biogas: Modeling and Simulations under Swedish Conditions |
| W1126637212 | FIALA_2015 | CARBON FOOTPRINT OF ELECTRICITY FROM ANAEROBIC DIGESTION PLANTS IN ITALY |
| W2312683967 | AZAPAGIC_2016 | Life Cycle Environmental Impacts of Electricity from Biogas Produced by Anaerobic Digestion |
| W2556832018 | KRÜMPEL_2016 | Demand-driven biogas production in anaerobic filters |
| W2592402297 | NELLES_2017 | Demand-driven biogas production by flexible feeding in full-scale – Process stability and flexibility potentials |
| W2724845225 | BOZZETTO_2017 | Greenhouse gas emissions of electricity and biomethane produced using the Biogasdoneright™ system: four case studies from Italy |
| W2768561067 | DALE_2017 | Sequential crops for food, energy, and economic development in rural areas: the case of Sicily |
| W2805047418 | LIAO_2018 | Anaerobic co-digestion of multiple agricultural residues to enhance biogas production in southern Italy |
| W2794973208 | THRÄN_2018 | Flexible Biogas in Future Energy Systems—Sleeping Beauty for a Cheaper Power Generation |
| W2888708623 | LIAO_2018 | Spatial analysis of feedstock supply and logistics to establish regional biogas power generation: A case study in the region of Sicily |
| W2790521460 | PORTO_2018 | A GIS‐based spatial index of feedstock‐mixture availability for anaerobic co‐digestion of Mediterranean by‐products and agricultural residues |
| W2914822267 | PROCHNOW_2019 | The Future Agricultural Biogas Plant in Germany: A Vision |
| W2988479878 | EUVERINK_2019 | Rambling facets of manure-based biogas production in Europe: A briefing |
| W2969910225 | CLANCY_2019 | Promoting agricultural biogas and biomethane production: Lessons from cross-country studies |
| W2907370432 | HUISINGH_2019 | Investigating energy and environmental issues of agro-biogas derived energy systems: A comprehensive review of Life Cycle Assessments |
| W2941197762 | BLENGINI_2019 | Life Cycle Assessment of a Biogas-Fed Solid Oxide Fuel Cell (SOFC) Integrated in a Wastewater Treatment Plant |
| W3004153001 | PILARSKI_2020 | 15 Years of the Polish agricultural biogas plants: their history, current status, biogas potential and perspectives |
| W3102984777 | DACH_2020 | Biogas Plant Exploitation in a Middle-Sized Dairy Farm in Poland: Energetic and Economic Aspects |
| W4205695009 | MAZURKIEWICZ_2022 | Energy and Economic Balance between Manure Stored and Used as a Substrate for Biogas Production |
| W4309852604 | MAZURKIEWICZ_2022 | Analysis of the Energy and Material Use of Manure as a Fertilizer or Substrate for Biogas Production during the Energy Crisis |
| W4386850198 | MAZURKIEWICZ_2023 | Loss of Energy and Economic Potential of a Biogas Plant Fed with Cow Manure due to Storage Time |
| W4321781369 | DACH_2023 | Reduction of Greenhouse Gas Emissions by Replacing Fertilizers with Digestate |
| W4386913257 | MAZURKIEWICZ_2023 | The Impact of Manure Use for Energy Purposes on the Economic Balance of a Dairy Farm |
| W4405283529 | KUSZ_2024 | The Capacity of Power of Biogas Plants and Their Technical Efficiency: A Case Study of Poland |
Topic modeling (STM)
Detect topics within a group using Structural Topic Modeling, creating sub-groups based on linguistic similarities.
# Prepare STM data (~30 min)
groups_stm_prepare <- birddog::sniff_groups_stm_prepare(
groups,
group_to_stm = "c1g3"
)17 topics is the best fit.
groups_stm_prepare$plots[['metrics_by_k']]
groups_stm_prepare$plots[['exclusivity_vs_coherence']]
# Run STM (~35 sec)
groups_stm_run <- birddog::sniff_groups_stm_run(
groups_stm_prepare,
k_topics = 17,
n_top_documents = 20
)
groups_stm_run$topic_proportion |>
dplyr::mutate(topic_proportion = round(topic_proportion, 3)) |>
gt::gt()
groups_stm_run$top_documents |>
dplyr::group_by(topic) |>
dplyr::arrange(dplyr::desc(gamma)) |>
dplyr::slice_head(n = 3) |>
dplyr::select(-DI) |>
gt::gt() |>
gt::text_transform(
locations = gt::cells_body(columns = document),
fn = function(x) {
glue::glue('<a href="https://openalex.org/{x}" target="_blank">{x}</a>')
}
)Session info
sessionInfo()
#> R version 4.5.2 (2025-10-31)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Manjaro Linux
#>
#> Matrix products: default
#> BLAS: /usr/lib/libblas.so.3.12.0
#> LAPACK: /usr/lib/liblapack.so.3.12.0 LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
#> [3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
#> [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
#> [7] LC_PAPER=en_US.UTF-8 LC_NAME=C
#> [9] LC_ADDRESS=C LC_TELEPHONE=C
#> [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: America/Cuiaba
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] birddog_1.0.5
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.56 bslib_0.10.0
#> [4] ggplot2_4.0.2 htmlwidgets_1.6.4 ggrepel_0.9.6
#> [7] lattice_0.22-7 vctrs_0.7.1 tools_4.5.2
#> [10] generics_0.1.4 tibble_3.3.1 janeaustenr_1.0.0
#> [13] tokenizers_0.3.0 pkgconfig_2.0.3 Matrix_1.7-4
#> [16] data.table_1.18.2.1 RColorBrewer_1.1-3 S7_0.2.1
#> [19] desc_1.4.3 gt_1.3.0 lifecycle_1.0.5
#> [22] compiler_4.5.2 farver_2.1.2 stringr_1.6.0
#> [25] textshaping_1.0.4 ggforce_0.5.0 graphlayouts_1.2.2
#> [28] litedown_0.8 openalexR_2.0.2 SnowballC_0.7.1
#> [31] htmltools_0.5.9 sass_0.4.10 tidytext_0.4.3
#> [34] yaml_2.3.12 lazyeval_0.2.2 plotly_4.12.0
#> [37] pillar_1.11.1 pkgdown_2.2.0 jquerylib_0.1.4
#> [40] tidyr_1.3.1 MASS_7.3-65 cachem_1.1.0
#> [43] viridis_0.6.5 commonmark_2.0.0 tidyselect_1.2.1
#> [46] digest_0.6.39 stringi_1.8.7 dplyr_1.2.0
#> [49] purrr_1.2.1 polyclip_1.10-7 fastmap_1.2.0
#> [52] grid_4.5.2 cli_3.6.5 magrittr_2.0.4
#> [55] dichromat_2.0-0.1 ggraph_2.2.2 tidygraph_1.3.1
#> [58] utf8_1.2.6 withr_3.0.2 scales_1.4.0
#> [61] rmarkdown_2.30 httr_1.4.7 igraph_2.2.1
#> [64] otel_0.2.0 gridExtra_2.3 ragg_1.5.0
#> [67] memoise_2.0.1 evaluate_1.0.5 knitr_1.51
#> [70] viridisLite_0.4.3 markdown_2.0 rlang_1.1.7
#> [73] Rcpp_1.1.1 glue_1.8.0 tweenr_2.0.3
#> [76] xml2_1.5.2 jsonlite_2.0.0 R6_2.6.1
#> [79] systemfonts_1.3.1 fs_1.6.7